Research Overview
The Liptak group elucidates the mechanisms of bioinorganic enzymes to aid the development of novel antibiotics and catalysts for biomedical and environmental applications.
Currently, we are particularly focused on enzymes involved in metal tetrapyrrole biosynthesis and degradation.
Several pathogenic bacteria employ heme iron acquisition pathways to procure iron duing infection, and we are examining the mechanisms of the heme degrading enzymes from this pathway to aid antibiotic development.
We are also investigating the mechanisms of class II chelatases, which catalyze metal insertion during metal tetrapyrrole biosynthesis, with the aim of designing synthetic enzymes for the biosynthesis of non-natural metal tetrapyrroles for alternative energy applications.
Finally, we have found important applications of our approaches in the field of molecular photophysics.
A common challenge in all three of these research areas is understanding the electronic structure.
For most proteins and main group complexes, reactivity and properties can be predicted and understood solely based upon geometric structure.
In these species, geometric structure determines electronic structure which defines reactivity and properties.
However, for transition metal-containing proteins and complexes, a single geometric structure can yield a broad range of oxidation, spin, and configurational states based upon subtle structural changes.
Each of these unique electronic structures has unique reactivity.
Similarly, an understanding of molecular photophysics requires insight into excited state electronic structure, which also does not obviously follow from the ground state geometry.
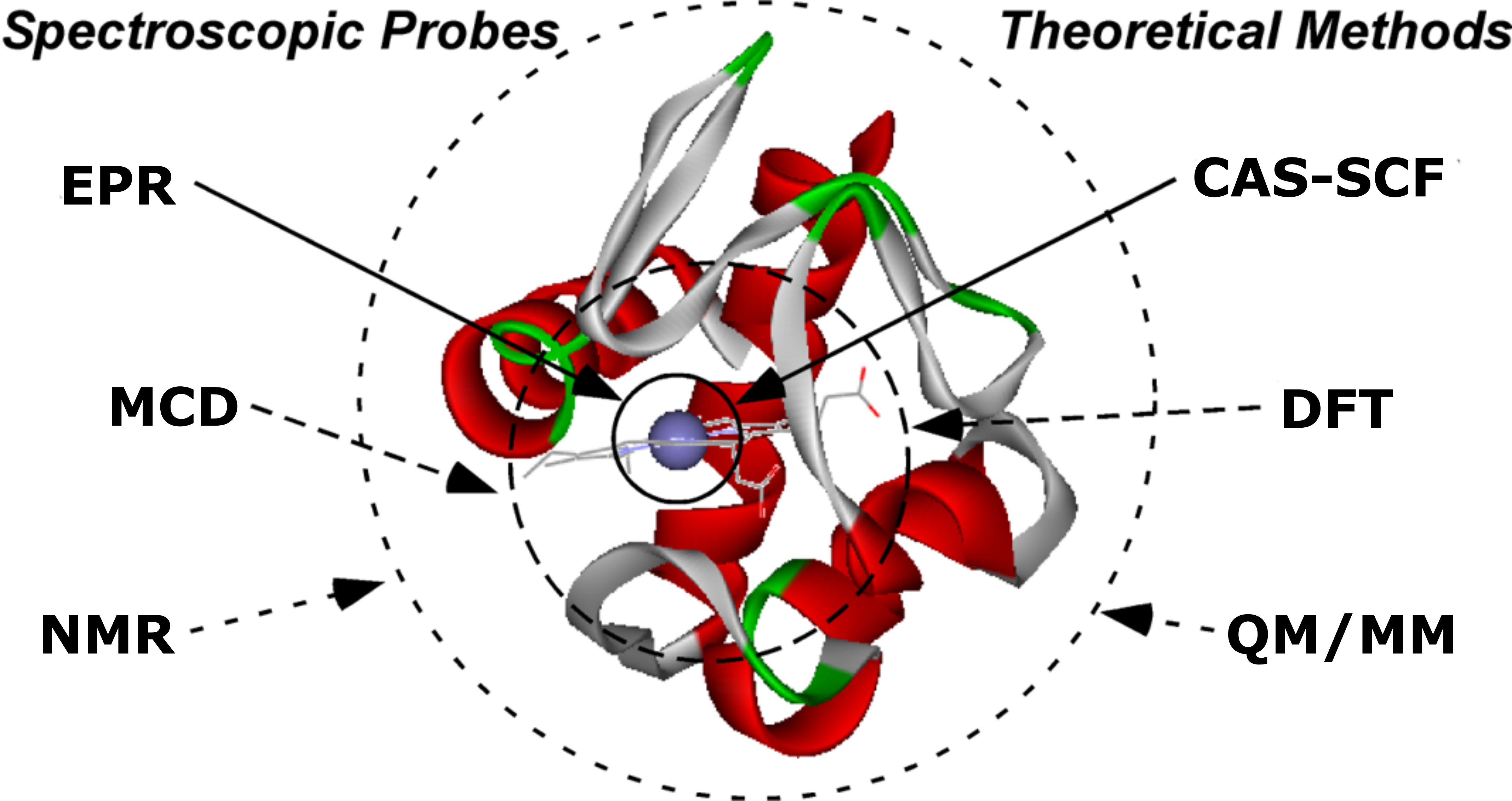
To achieve these goals, we employ biochemical, spectroscopic, and computational approaches.
Biological samples are prepared via recombinant protein expression, and key reactive intermediates are isolated using oxygen-free techniques.
These intermediates, and their analogues, are characterized using a variety of spectroscopies, including: magentic circular dichroism (MCD), nuclear magnetic resonance (NMR), and electron paramagnetic resonance (EPR).
To aid interpretation of our spectroscopic data, we model these species using quantum mechanical calculations.
 Metal Tetrapyrrole Degradation
Metal Tetrapyrrole Degradation
The longest-running project in the Liptak group is our study of
Staphylococcus aureus IsdG and
Mycobacterium tuberculosis MhuD.
These heme oxygenases are part of the heme iron acquisition pathways these two pathogens employ to harvest iron during human infection.
As these bacteria require micromolar iron to support their growth, both pathways are promising targets for novel antibiotic treatments.
Humans employ heme oxygenases to recycle iron from heme, and it is critical to identify unique properties of the bacterial heme oxygenases in order to specifically target the bacterial enzymes.
Along these lines, our focus has been to elucidate the novel enzymatic mechanisms of IsdG and MhuD.
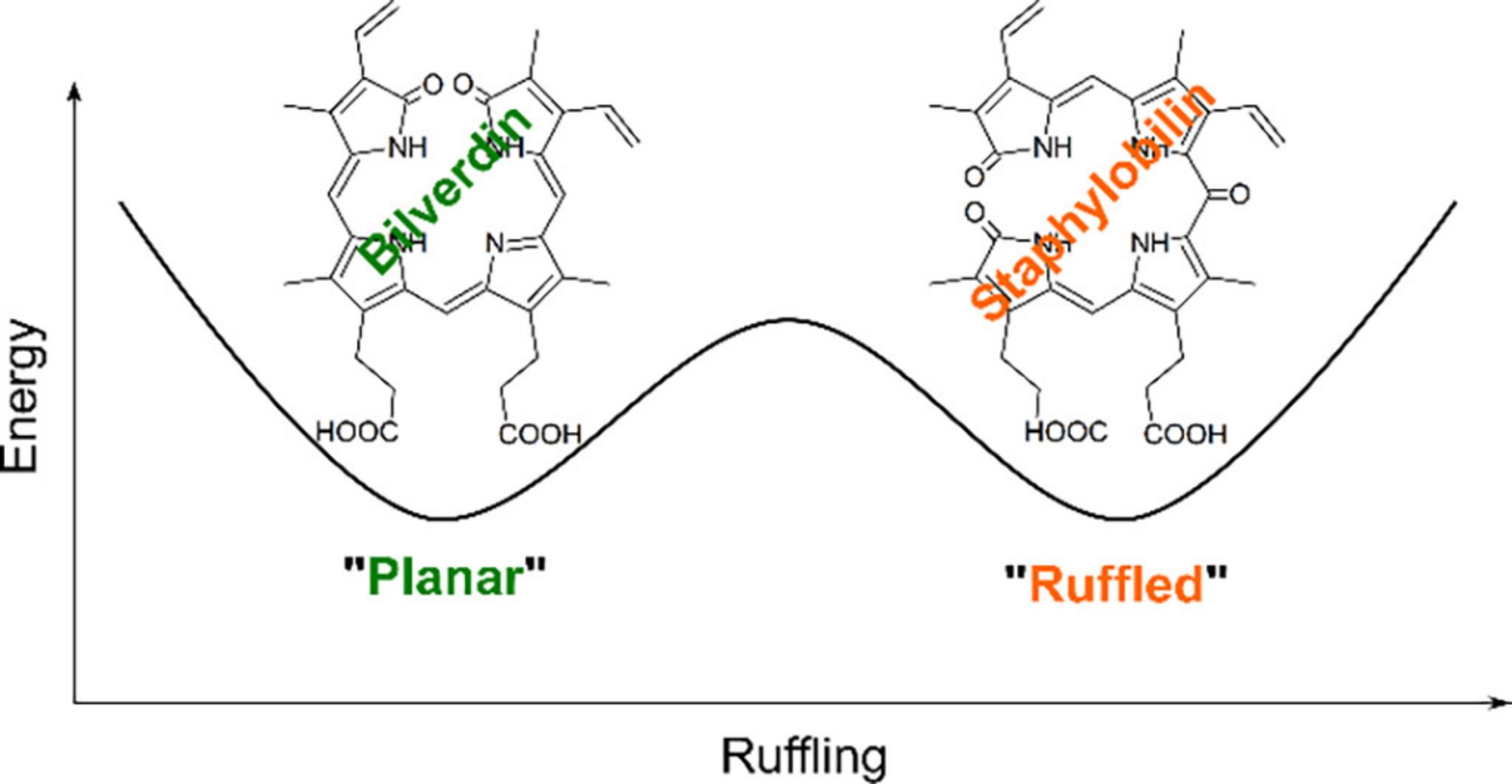
One of our most exciting discoveries regarding IsdG and MhuD is that both enzymes bind a dynamic heme substrate.
Two heme conformations, one being planar and the other exhibiting an out-of-plane "ruffled" porphyrin, are in dynamic equilibrium within the enzyme active site.
Surprisingly, the two heme conformations have distinct electronic structures: a
2Eg electronic ground state for planar heme and a
2B2g ground state for "ruffled" heme.
Technically, this is a hidden pseudo Jahn-Teller distortion.
But, what is most exciting, is that enzyme-catalyzed oxygenation of the two conformations yields two distinct enzymes products.
 Metal Tetrapyrrole Biosynthesis
Metal Tetrapyrrole Biosynthesis
A newer project in the Liptak group is concerned with transition metal insertion during metal tetrapyrrole biosynthesis.
Metal tetrapyrroles are diverse scaffolds that chelate a transition metal within a square planar coordination geometry.
Nature uses magnesium(chlorophyll), iron(heme), cobalt(vitamin B
12) and nickel (cofactor F
430) to catalyze a wide-range of reactions, such as: light harvesting, oxygen activation, dehalogenation, and methanogenesis with earth abundant metals.
In principle, non-natural metal tetrapyrrole combinations could be used to catalyze a variety of reactions critical for next generation energy applications with earth abundant metals.
However, a critical challenge is our limited understanding of metal selectivity by chelatase enzymes.
Our group is working to clarify the origin(s) of metal selectivity by chelatases.
To date, our research has focused on two "ancestral" chelatase enzymes (
Archaeoglobus fulgidus CbiX
S and
Methanosarcina acetivorans CfbA.
These enzymes are approximately half the size of the chelatases that insert iron (HemH) and cobalt (CbiK) during heme and vitamin B
12 biosynthesis, respectively.
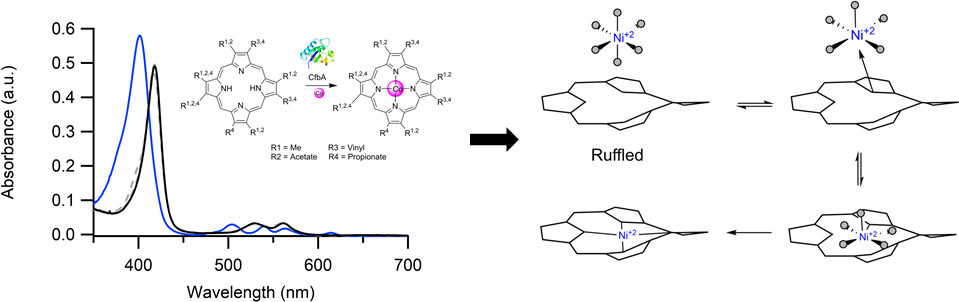
CfbA catalyzes nickel insertion during cofactor F
430 biosynthesis.
we have discovered that CbiX
S inserts a labile Ni(II) into a "ruffled" tetrapyrrole.
This lability strongly suggests the metal selectivity is under thermodynamic control.
 Molecular Photophysics
Molecular Photophysics
Finally, we have found important applications of our combined spectroscopic and computational approach in the field of molecular photophysics.
Organic fluorophores have great promise for next-generation lighting technologies, but it will be important for researchers to develop exquisite control over the photophysical properties of these species.
Emission wavelength, emission bandwidth, and brightness are three particularly important properties.
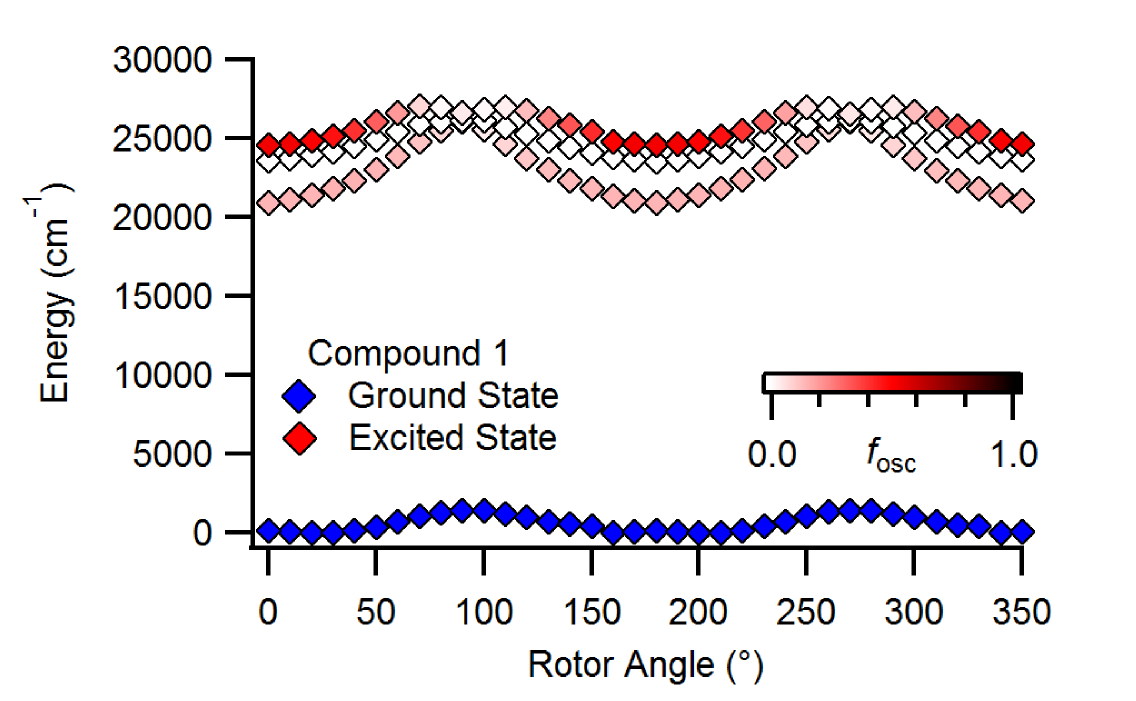
All of these properties are derived from the details of the ground and excited state potential energy surface, which itself depends upon the molecular environment.
Our group has worked to unveil photophysical mechanisms and predict photophysical properties.
Most notably, we discovered a novel photophysical mechanism responsible for the aggregation-induced emission (AIE) of hydrazone-based fluorophores.
Boron difluorohydrazone fluorophores exhibit viscosity-dependent emission, but not polarity-dependent emission, which means that their AIE does not arise from the common twisted-intramolecular charge transfer (TICT) mechanism.
Computational modelling led to the proposal of the suppression of Kasha's rule (SOKR) mechanism where access to a molecular conformation that promotes internal conversion is suppressed.
More recently, we have identified the structural origin of the "giga" Stokes shifts observed in triazolopyridinium-based fluorophores.
 Job Openings
Job Openings
We currently have an opening for a
graduate student interested in the
Metal Tetrapyrrole Degradation project.
Please contact Prof. Liptak for more information (
mliptak@uvm.edu).







